Last week, I took a look at the very illogical excuse that virologists make in regard to why they are unable to purify and isolate the particles that are claimed to be “viruses” directly from the fluids of a sick human or animal. As a reminder, below is the response I received from biologist Thomas Baldwin, who studies “pathogenic” plant “viruses” and goes by the Twitter handle Sense_Strand:
It is claimed that there are just not enough of these “viral” particles within the fluids and thus, the purification procedures will result in too little of the “virus” remaining after these steps are performed. Due to this lack of particles, it is claimed that the “viruses” can not be found in electron microscopy images, and it is for this very reason that the “virus” particles must be grown in cell culture so that the “virus” can replicate to a large enough number in order to be visualized and studied. While I won’t rehash my counterargument here, I will allow Debunked to help me demonstrate why this is a ridiculous excuse:
When virologists claim incredible numbers like that, it is pretty reasonable to conclude that there should be plenty of “viral” particles within the fluids of a sick animal or human in order to purify, isolate, visualize, characterize, and study. Alas, virologists defiantly cling to their laughable excuse in order to cover up for the fact that they just cannot find the assumed “viral” particles anywhere directly within the fluids. While this statement clearly defies logic, the lack of “virus” is only one aspect of the excuse. There is another component that is used to explain why, even if they could purify and isolate the particles, it wouldn’t ultimately matter. Beyond the lack of enough “viral” particles within the fluids, virologists claim that there are not enough “infectious” particles present after purification in order to be able to “infect” an animal or human on order to prove pathogenicity. It is stated that this purification process damages the “virus” and causes it to lose “infectivity.” This excuse was illustrated in a response interviewer Djamel Tahi received from HIV “discoverer” Luc Montagnier:
“I believe we published in Science (May 1983) a gradient which showed that the RT had exactly the density of 1.16. So one had a ‘peak’ which was RT. So one has fulfilled this criterion for purification. But to pass it on serially is difficult because when you put the material in purification, into a gradient, retroviruses are very fragile, so they break each other and greatly lose their infectivity.”
“I repeat we did not purify. We purified to characterise the density of the RT, which was soundly that of a retrovirus. But we didn’t take the “peak”…or it didn’t work…because if you purify, you damage. So for infectious particles it is better to not touch them too much.”
https://viroliegy.com/2022/02/13/montagniers-monster/
As can be seen, if the particles are purified, it is assumed that they lose their “infectivity.” Thus, virologists must not touch their fragile little “virus” particles too much or they will be damaged and will not work properly. With statements like this, it makes the story about how these non-living entities somehow survive the harsh environmental conditions of the great outdoors in order to invade a body, bypass the hosts “immune system,” and hijack the cells so that it can create more copies of itself, seem rather ridiculous. According to virologists, in order to retain “infectivity,” the “virus” particles must remain unpurified and proceed to be mixed into a foreign animal or cancer cell with toxic antibiotics, antifungaks, fetal calf blood, chemicals and “nutrients,” etc. and incubated for days. However, this is normally not enough to create the necessary “infective” particles, so virologists will remove the top layer of one culture and then add it to another culture with a fresh round of toxic compounds mixed in. This new culture is then incubated further until signs of cell death are observed. Only then can there be enough “infectious viral” particles to visualize and establish pathogenicity.
To the outside observer who looks at this critically and logically, it is clear that all virologists are doing is creating a toxic soup of many foreign and chemical elements in which they get to claim a “virus” resides within. This sludge is then forcefully and unnaturally inoculated into animals in many disgusting ways, either through the nose, the skin, the muscles, the eyes, the throat, the stomach, the brain, or even the testicles. Virologist then determine what is an “infectious” dose based upon how much of this toxic soup is used as an injection into the animal at the time any symptoms appear. Virologists will determine how much “virus” is present in the soup by utilizing either one of two methods: the tissue culture infectious dose (TCID50) and the plaque assay. Let's examine these methods briefly and then see, according to virology's own theoretical narrative, how many “viral” particles it takes to cause infection and disease. We can then determine whether or not it is reasonable to believe that there are not enough “infectious” particles present after purification and isolation in order to determine pathogenicity.
Tissue Culture Infectious Dose (TCID50)
This first method for estimating how many “virus” particles are necessary for “infection” relies on the observation of the cytopathogenic effect (CPE) that is created during the cell culture experiment. CPE is an effect that is observed when the cells start to die and break apart during the cell culture process after the cell has been starved and poisoned. To calculate how many “viruses” they believe are present and “infectious,” virologists will use varying “virus” dilutions that are added as an endpoint dilution to host cell populations in a 96 well plate format. They will then incubate these mixtures until a cytopathic effect can be observed. The wells are either inspected by visually counting the CPE in the affected wells or by using assay readouts. Once 3 of the same CPE readings in separate cells for the same dilution are observed, the dose is calculated using one of various mathematical equations. The dilution at which 50% of the cell cultures are “infected” is determined and used to mathematically calculate a TCID50 result:
Tissue Culture Infectious Dose (TCID50) Assays: How to determine virus infectivity?
TCID50 assays: How do they work?
“50% Tissue Culture Infectious Dose (TCID50) assays are virus titration experiments which can be used to quantify virus titers by investigating the cytopathic effects of a virus on an inoculated host cell culture4. Compared to the widely used plaque assays, which are also used in virus quantification, TCID50 assays offer the advantage that even viruses that do not form plaques or infect cell monolayers can be quantified.
In TCID50 assays, varying virus dilutions are added as an endpoint dilution to host cell populations with the same number of cells and incubated until a cytopathic effect can be seen. Here, the TCID50 value represents the amount of virus dilution required to induce cytopathic effects in 50% of wells containing the inoculated cell culture after a defined period of time.
TCID50 assays assess this threshold either by visually counting the number of affected wells or by using cell viability assays as readout. The TCID50value is determined when the cytopathic effect or cell viability assay read-out appear the same for a dilution in 3 separate readings. An example of the application of cell viability/toxicity assays for the evaluation of viral cytopathic effects can be found in the AN 363: Viral cytopathic effects measured in a drug discovery screen.
TCID50 calculation
The results of 50% Tissue Culture Infectious Dose (TCID50) assays can be analysed by different calculations 5. Several mathematical approaches have been developed for this purpose, including the Reed-Muench 4, Spearman-Kärber or Weil method. The formula after Reed-Muench is depicted as an example below.
Where I is the interpolated value of the 50% endpoint and h is the dilution factor.
Since most often, the exact 50% endpoint is not observed in TCID50 assays, an approximate value can be obtained factoring in the dilutions closest below and above the 50% threshold. Independent of the method, the dilution at which 50% of the cell cultures are infected is determined and used to mathematically calculate a TCID50 result which is expressed as 50% infectious dose (ID50) per millilitre (ID50/mL) after a defined period of time. For example, if 0.2 mL of a 1:10,000 virus dilution infects 50% of the cells in 2 days the titer is expressed as 104 TCID50/0.2 mL in 2 days.”
As can be seen, this method relies on the observation of CPE as evidence of a “virus” and then attempts to calculate how many of these invisible entities reside within the fluids. However, as should be expected when dealing with attempts to count something that can not be seen, this method has its drawbacks. For one, the Poisson distribution that is utilized, which takes the TCID50 value and multiplies it by 0.7, is admittedly only an approximation and is said to not always be true. The serial dilution method itself is also a source of error by its very nature. If any fluid remains at the end of the pipette used to suction out the “virus,” it is said that this can greatly influence the quantification results. Another issue is attempting to keep all of the variables exactly the same across all cultures, which is admitted to not always be the case. Thus, there is a lot of guesswork and assumption involved in calculating the infectious dose of the unseen entity:
Timeless TCID50: One solution to many viruses
From dilutions to titres
“TCID50 values give an indication of how many viruses is needed to have CPE in 50% of the cells. But how to go from this to the actual amount of virus per ml? The formula is quite simple, and it consists in multiplying the TCID50 value by 0.7. This comes from the Poisson distribution applied to viral infection which states that, in a fully permissive cell line, the probability of reaching 50% infection is achieved by a multiplicity of infection of approximately 0.7. This is not always true, but it’s a good approximation for most applications.
The troubles of counting viruses
As accurate as one can be, counting viruses is never easy. First, serial dilutions are -by their own nature- a source of error. Second -and this is particularly relevant for high titres of virus- even the tiniest volume that remains attached to the very end of a pipette tip can carry enough viral particles to make a substantial difference in the quantification. Third, the biological variation of the system is high. Plate the same amount of cells, add the same amount of virus, stop the infection at the same time, and the percentage of infection may be close, but never exactly the same.
Finally, when assessing a treatment that (as you would hope!) decreases virus titres, the amount of virus may fall below the assay detection threshold.”
https://virologyresearchservices.com/2019/03/29/timeless-tcid50-one-solution-to-many-viruses/
Plaque Assays
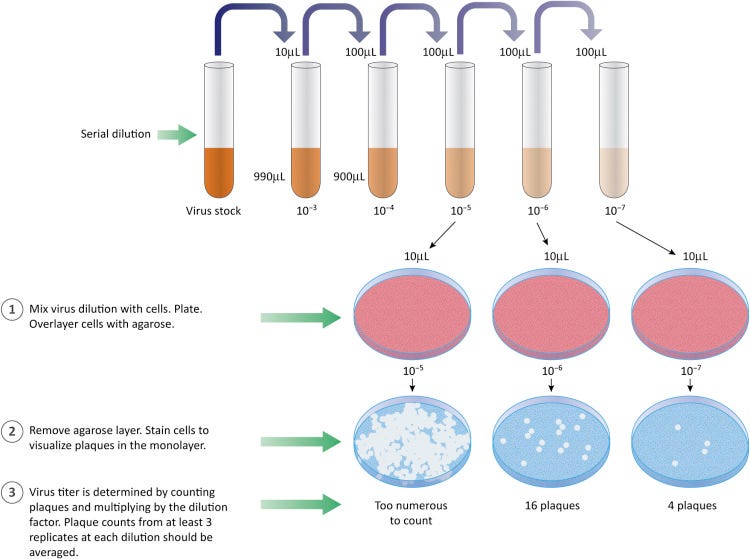
However, if relying on an indirect effect and spotty mathematical equations to calculate how many “viruses” it takes to “infect” a cell is not to your liking, you may appreciate this next method even less. Plaque assays also rely on the observation of CPE in cultured cells. As the cells break apart and die, the “viral” particles are assumed to travel to neighboring cells, infecting them and creating plaques, or holes, in the dish. The cells are then fixed and stained, killing everything in order to be observed. It is said that the cells that remain adhered to the surface are assumed to be uninfected, and any observed plaques are assumed to arise from cell death caused by “viral infection.” Virologists will look for the dilution that led to the optimal observed plaques; too little dilution leads to too many plaques while too much dilution yields none. The titre is then calculated using arithmetic based on the volume of the aliquot added to the cells and the sample dilution the aliquot was drawn from. The assay is designed so that each plaque represents infection by only a single “viral” particle:
Measuring infectious virus: the plaque assay
Infection and plaque formation
Plaque assays require cultured cells susceptible to infection by the virus of interest. The cells are first seeded onto a surface they can adhere to and grow on, then left overnight to form a confluent monolayer (a cohesive sheet of cells covering the entire growth surface). A virus sample is then diluted several times, and an aliquot of each dilution is added to a dish or well of cells. An incubation period allows the virus to attach to target cells before removing the inoculum. The culture is then covered with a medium containing nutrients and a substance, such as agarose or methylcellulose, forming a gel or semisolid overlay. Infectious virus particles that enter cells and replicate can then trigger the release of progeny virions. The gel restricts particle movement so that newly produced viruses can only infect neighbouring cells. If the virus kills infected cells, the dead (or dying) cells detach and create a hole in the monolayer through lysis or other means. This space – now devoid of cells –is called a plaque and appears as circular spots on the growth surface.
The plaques are allowed to grow until visible to the naked eye. The cells are then fixed with formaldehyde to lock cellular structures while killing the cells and virus. Dyes that stain cells are added for contrast, making plaques easier to see. Purple violet stains the cells purple, while plaques, lacking cells, remain clear. Cells that remain adhered to the surface are assumed to be uninfected, and apparent plaques are assumed to arise from cell death caused by infection. That is why the virus dilutions must be added to confluent monolayers with no gaps that might later be mistaken for plaques.
Viral titre: PFU/ml
Multiple dilutions of the stock sample are analysed to identify one or more dilutions that give rise to a countable number of plaques. At the lowest dilutions, too many infectious particles will destroy large swaths of the cell monolayer or create plaques too numerous and overlapping to distinguish. At the highest dilutions, there may be no plaques at all. At the optimal dilutions, plaques are counted to determine the titre of the original stock sample, typically reported as the number of plaque-forming units per millilitre (PFU/ml).
For a given plaque count, the stock titre can be calculated by simple arithmetic based on the volume of the aliquot added to the cells and the sample dilution the aliquot was drawn from. As a basic example, if 35 plaques were counted when a 0.1 ml aliquot of the 10-5 dilution was added to the cells, the titre of the undiluted stock is 3.5×107 PFU/ml. For reliable titres, each sample dilution should be plated multiple times, at least in duplicate and preferably in triplicate. Furthermore, multiple dilutions may result in countable plaques. More elaborate formulas incorporating all relevant plaque counts are typically used to calculate titres.
PFU/ml vs IU/ml
The assay is designed so that each plaque results from infection by multiplying a single infectious virus particle. As such, PFU/ml is considered a measure of the number of infectious units per millilitre (IU/ml), with the caveat that one cannot be certain of a one-to-one ratio of plaques to infectious particles in the applied aliquot. Also, be aware that the titre of a sample is specific to the assay conditions used to determine it, as infectivity is influenced by many factors, such as the type of host cell, pH, and culture medium. Titres can differ by several orders of magnitude by changing key assay parameters.”
https://virologyresearchservices.com/2022/08/10/the-plaque-assay/
As with the TCID50, there are some definite drawbacks when trying to “accurately” estimate how much of an invisible entity can cause infection and disease using the plaque assay. For starters, as noted above, the “infectivity” is said to be influenced by many factors within the culture itself. This includes the type of host cell, the pH level, as well as the culture medium used. Thus, the calculated titres can vary wildly by several orders of magnitude just by changing the assay parameters. The determination of what exactly constitutes a plaque is very subjective as well, which can mislead the results. In other words, plaque counting is prone to human error.
Beyond these issues, as noted before, both TCID50 and plaque assays rely on the observation of a cytopathogenic effect in order to claim the presence and infectivity of any “virus.” However, CPE is not specific to “viruses” as there are many known factors that can result in this effect being observed that do not require the presence of a fictional entity in order to explain, such as:
Bacteria
Parasites
Amoebas
Chemical Contaminants
Age of the Cell
Incubation Temperature
Length of Incubation
Antibiotics/Antifungals
It is entirely unscientific to rely on an effect in order to presume a cause. However, this pseudoscientific concept is central to the cell culture method as well as to the processes utilized to guesstimate how many “infectious viruses” are present. In other words, there is no direct evidence any “viruses” are present in any sample used to determine an “infectious” dose. All of these results claiming how much “virus” is present and can ultimately cause an infection are entirely hypothetical and calculated based on the presence of cell death. It is nothing but guesswork.
However, if it wasn't clear that these numbers are pseudoscientific fraud, a 2003 OSHA White Paper looking at the determination of the infectious dose (ID) may help to sell the fact that virologists honestly have no clue what an infectious dose is. While this refers to calculating ID using animals, the same criticisms can apply to utilizing lab-created cell cultures as a surrogate. The White Paper concluded that there is no clear definition of what an infectious dose is and that there is no single standardized method for determining ID. The extrapolation of data to humans is unreliable and is a poor surrogate for human responses. There are various secondary interactions that can impact the estimates. The “pathogens” vary wildly in “virulence” and data on the ID via route of exposure is unavailable. In other words, virologists just make things up as they go along:
OSHA Infectious Dose White Paper
“In summary, the studies described above support ABSA's position that attempts to develop quantitative values for human infectious dose are not currently feasible. Infectious dose values developed using past studies would not accurately characterize the relative hazard of pathogenic organisms in humans. The reasons for this conclusion are:
• Lack of a clear and universally acceptable definition of the term "infectious dose."
• There is no single standardized protocol for testing infectious dose in animals, making legitimate controlled comparisons ofstudy results very difficult.
• Extrapolation of infection and toxicity data among animal species and from animals to humans has proven to be unreliable for most biological (and chemical) agents.
• Inbred animal strains are a poor surrogate for predicting human response, as humans are a highly variable outbred population.
• Infectious dose is affected by numerous, complex secondary interactions to include condition of the host, its genetics, and previous exposure to the biological agent or vaccine. Risk estimates must take these and many other factors into consideration.
• Bacteria of a single species can vary widely in virulence and infectious dose. It is not possible to make a broad or generalized statement about the infectious dose of a species of bacteria.
• Infectious dose in part depends on the route of exposure. A complete picture of a single pathogen's infectious dose profile requires inhalation, percutaneous, oral, im, ip, iv, etc. data. These data are currently unavailable.”
Now that we know that this process for counting the invisible “viruses” is fraught with drawbacks that leave the “accuracy” of these results extremely questionable, let's take a look at some of these wildly varying estimates in order to see how many particles are necessary to cause an infection according to virology's fictional narrative. As “SARS-COV-2” is the soup du jour at the moment, here are a few key highlights from a systematic review of many studies attempting to find the minimal ID for the “novel coronavirus.” What you will see from the August 2022 review is that there is absolutely no experimental data for humans, with one study admittedly presenting a hypothetical estimate (technically, they are all hypothetical). The minimal ID for “SARS-COV-2” is extrapolated from animal studies with estimates that vary wildly between the papers:
Minimum infective dose of severe acute respiratory syndrome coronavirus 2 based on the current evidence: A systematic review
“The main methods for reporting the infective dose were through tissue culture infectious dose (TCID50) and by counting plaque-forming units (PFU).33
In TCID50, the viral dose in 5% of inoculated tissue culture made pathological changes or cell death. PFU is estimated of viral concentration in plaque-forming units by measuring the number of particles that form a plaque.34 The minimum infective doses have been summarized in Table 2.
Human studies on infective dose of SARS-CoV-2
“We found no experimental studies that assess the infective dose in human, so we included observational human studies.”
“Finding the minimum infective dose of the virus can be extremely useful in determining the transmission pattern. This represents itself in inconsistent results across the included studies; similar viral load did not cause the same outcome. This indicates that despite having a similar minimum infective dose, the infection rate could differ so this minimum is not the same across the same population. On the other hand, there are some human studies which have shown some hypothetical infective viral dosages.”
Conclusion
“The results of this review suggest that one of the key factors to control the pandemic could be the study of virus transmission. The minimum infective dose is one of the main components of virus transmission. In this study, we have presented a range of minimum infective doses in humans and various animal species, yet such numbers can possibly vary between the individuals based on numerous factors. Measuring the minimum infective dose can provide a clearer overall understanding of the disease and its transmissibility and help better halt its spreading.”
https://journals.sagepub.com/doi/full/10.1177/20503121221115053#table2-20503121221115053
While it is fun to see how much their guesstimates can range between studies, the above paper doesn't give us a great idea as to what this minimal ID is for “SARS-COV-2” in terms of an actual number of particles it may take in order to cause infection. Let's see if we can put a number to it based upon what the “experts” are saying:
SARS-CoV-2 Infectious Dose
“Some experts estimate that exposure to as few as 1000 SARS-CoV-2 viral particles can cause an infection. This dose of virus could occur by inhaling 1000 infectious viral particles in a single breath, 100 viral particles in 10 breaths, or 10 viral particles in 100 breaths.”
https://www.clinlabnavigator.com/sars-cov-2-infectious-dose.html
According to the “experts,” as little as 1000 “SARS-COV-2” particles are enough to cause infection. Where did they get this magical number from? Who knows? However, a separate study gave an even smaller estimate of just 100 “viral” particles:
Review of infective dose, routes of transmission and outcome of COVID-19 caused by the SARS-COV-2: comparison with other respiratory viruses
“An accurate quantitative estimate of the infective dose of SARS-CoV-2 in humans is not currently feasible and needs further research. Our review suggests that it is small, perhaps about 100 particles.”
It appears that virologists have the ability to just pull any ID number that they want out of thin air. When we factor in their other imaginary numbers, such as those who are at “peak viral infection” harboring 10 to 100 billion “viral” particles while breathing out 10 million “viruses” per breath, it seems rather illogical to claim that there would not be enough “infectious virus” after purification in order to prove pathogenicity.
Examining the issue a bit further, let's look at a few other sources and see just how little “virus” is said to actually be necessary in order to cause an infection. According to virology's pseudoscientific narrative, would you believe that just one airborne particle can cause you to become infected? That is exactly what we are told by this next source which dealt a fatal blow to mask supporters everywhere. The researchers based their claim upon theoretical modeling (aren't they all), and concluded that only one airborne particle is enough to cause infection and disease:
What if just one airborne particle was enough to infect you?
“For some diseases, exposure to just a single airborne particle containing virus, bacteria or fungi can be infectious. When this happens, understanding and predicting airborne disease spread can be a whole lot easier.
That’s the result of a new study by a Lawrence Livermore National Laboratory (LLNL) scientist who developed a new theory of airborne infectious disease spread. This research, which appears in the journal Applied and Environmental Microbiology, demonstrated good agreement with data from Q fever, Legionnaire’s disease and Valley fever outbreaks. The authors hope to use it to understand and mitigate COVID-19 spread.”
https://www.llnl.gov/news/what-if-just-one-airborne-particle-was-enough-infect-you
Unfortunately, the authors did not give us any idea as to how many “viral” particles would be in one airborne particle. Hypothetically, it could be one “virion” or it could be quite a few more. Let's see if we can get some concrete guesstimates as to how many of the “viral” particles it may take to cause infection and disease. According to the CDC, “norovirus” only requires a few particles in order to cause infection and disease:
About Norovirus
“People with norovirus illness can shed billions of norovirus particles. And only a few virus particles can make other people sick.”
https://www.cdc.gov/norovirus/about/index.html
This works out to as little as 10 “viral” particles per the European CDC:
“Noroviruses are highly contagious and 10-100 viral particles may be sufficient to infect an individual.”
https://www.ecdc.europa.eu/en/norovirus-infection/facts
When looking to insect “viruses,” researchers set up an experiment with two “marked virus variants.” They exposed a population of caterpillars to both variants, and based upon these results, established a probability model to determine that it is theoretically possible for just one “virus” particle to cause infection and disease:
One Virus Particle Is Enough To Cause Infectious Disease
“Can exposure to a single virus particle lead to infection or disease? Until now, solid proof has been lacking. Experimental research with insect larvae has shown that one virus particle is theoretically enough to cause infection and subsequent disease.”
“Based on the assumption that every virus particle operates independently from all other virus particles, the researchers set up a probability model. This model predicts how many virus particles have caused an infection and how many different virus genotypes are present in infected hosts, such as plants, insects or people. The results of the infection experiment with the susceptible insects are in agreement with the model predictions. From this it can be derived that the virus particles have an independent effect, and that a single virus particle can indeed cause infection and/or disease.”
We have now officially gone from 1000 “viral” particles to just one particle. Let's see if any other sources make such a bold claim. In a CDC study that looked to establish a quantifiable estimation of how many “variola” (a.k.a. smallpox) particles are necessary to cause infection, the researchers concured that only one “virus” particle was sufficient to cause infection and disease:
The infectious dose of variola (smallpox) virus.
“Quantitative estimation of an individual's risk of infection due to airborne pathogens requires knowledge of the pathogen's infectious dose, in addition to estimates of the pathogen's airborne concentration and the person's exposure duration. Based on our review of the published literature on poxvirus infection, we conclude that the infectious dose of variola (smallpox) virus is likely one virus particle and that infection can be initiated in either the upper respiratory tract or pulmonary region. Studies of airborne transmission of poxvirus in monkeys and rabbits show that primary infection can occur in both regions of the respiratory tract. A quantitative study of poxvirus inhalation transmission in rabbits indicates that the deposition of one pock-forming unit (PFU) carried on respirable particles can cause infection. Findings in both in vitro and in vivo studies of the number of virus particles comprising a PFU are consistent with a "one-hit" phenomenon--namely, the cellular uptake of just one virus particle can lead to infection of a cell or an area of cell growth, creating a pock (an infected area of cells). Variability in virulence among different virus strains may involve differences in the probability of infection per virus particle, where a highly virulent strain has a probability close to one of successful infection for each virus particle.”
“On balance, we believe there is adequate in vitro and in vivo evidence that infection can be produced by a single particle of variola virus. Across different experimental systems the number of poxvirus per infectious unit has been found to vary, but it appears that favorable conditions enable all virus particles to infect (Overman & Tamm, 1956; Parker, Bronson, & Green, 1941; Sprunt & McDearman, 1940).”
https://www.cdc.gov/niosh/nioshtic-2/20037359.html
The CDC has also stated that only one “viral” particle from rabies is enough to cause infection and disease:
Rabies
“During clinical disease, millions of viral particles may be found intermittently in the saliva. In theory, only a single rabies particle or virion is required to result in a productive infection.”
https://www.cdc.gov/rabies/diagnosis/accuracy.html
This one “viral” particle is supported as well by OSHA in regard to the Ebola “virus:”
Ebola
“In areas of Africa where Ebola viruses are common, suspected reservoirs include primate and bat populations. While there are no known animal reservoirs of the disease in the U.S., there is concern related to possible spread of EVD among human populations due to the availability and reach of global travel. Under certain conditions, exposure to just one viral particle can result in development of EVD. Depending on the strain and the individual infected with the disease, EVD may be fatal in 50-90 percent of cases.1”
And finally, the measles “virus” is also considered so virulent that only one “viral” particle can cause infection and disease:
Measles
https://microbeonline.com/infective-dose-and-lethal-dose/
It is clear that, according to virology's own pseudoscientific narrative, only one “viral” particle is theoretically necessary in order to cause infection and disease. Therefore, there is absolutely no reason to assume that there are not enough infectious “viral” particles remaining after purification and isolation directly from the fluids to be used in order to prove pathogenicity. The reason this excuse is presented is because virologists are unable to recreate disease using just the fluids from a sick host. In order to even attempt to show pathogenicity, they claim that the unpurifued fluids must be added to a foreign animal or cancer cell along with antibiotics, antifungals, fetal calf blood, chemicals, “nutrients,” etc. and then injected unnaturally into an animal either through the nose, the blood, the throat, the skin, the brain, the stomach, the eyes, the feet, and/or the testicles. There is nothing natural or scientific about this process.
However, as virology is pseudoscience, it falls back on unfalsifiable concepts in order to excuse away the lack of scientific evidence. Instead of being able to find the “virus” particles directly in the fluids, they get to claim that there is not enough “virus” there despite their own numbers making this an impossibility. As virologists know that they can not prove pathogenicity using nothing but the assumed “viral” particles, they get to claim that the purification process creates a yield loss and that the “viruses” lose infectivity. Therefore, virologists get to excuse away that they can not show that the particles created after the cell culture experiment ever existed within the fluids of a sick host to begin with. They get to explain away that they can not prove pathogenicity using nothing but the assumed “viral” particles without culturing. Yet, despite their excuses, the numbers supplied by virologys own pseudoscientific narrative fail them. With hundreds of billions of “viruses” at peak infection, there is absolutely no reason that virologists should not be able to purify and isolate the assumed “viral” particles directly from the fluids of a sick human or animal. If only one “viral” particle is necessary theoretically in order to cause infection and disease, there is no reason virologists cannot use a purified sample to prove pathogenicity naturally via the scientific method. By their own admission, all it takes is just one particle to cause infection and disease. One particle from a sea of billions.
discussed the ridiculousness of the latest fear-mongering over tick-borne diseases. did an excellent follow-up on her expose on the faults with the peer review system. took a look at the dire state of science. teamed up with Dr's. Cowan and Kaufman to look at the rift developing in the freedom community.













What would you say if we were told that there are Extra Terrestrials living among us, once even one settles in a town there will soon be too many of them to count, and then they'll move on to the next town......But we cannot pick a bunch of people from any town, look through them, and find these creatures? That we have to take such a group of people, put them in a large hall with no windows or doors, place all sorts of things and living entities in the room, provide the people with minimal nutrition, spray the room with chemicals.... and then we'll see these invaders from outer space.
Well-written and concise, as always! Great job pointing out the lack of quantitative as well as qualitative evidence for what is causing "infection". Without properly purified and isolated particles, there is nothing to carefully study and characterize to name as the cause of dis-ease. Thank you for the link to my work as well, it is always appreciated.